Data in chemistry publications is very standardized which makes it possible (not easy) to think about robotic extraction of information. I’ve blogged earlier about the use of text, but what about graphics? This post shows the potential, but also the current unnecessary destruction of data. You don’t need to be a chemist to understand the issue.
Types of graphical object that occur frequently in chemistry are
- chemical structure diagrams (more later)
- graphs (i.e. plots, not topology, though these also occur)
- spectra (used to probe the nature of compounds and also to act as fingerprints.
Here I show some proton-NMR spectra (1H NMR), which are very powerful ways of looking into molecules containing hydrogen atoms (almost all do). It’s closely related to NMRI used for medical imaging. What is remarkable is its precision – the frequency used is (often) 500 Mhz (i.e. 5 * 108 per second. Because of this precision the frequency axis is usually expressed in parts per million (ppm). The scale runs from 10 to 0 ppm. This is recorded digitally, usually with 2N points, such as 8192, 16384 or even more. So that means that for each ppm there are about 1000 points or more.
The values and the precise shapes of the peaks are very important. They are usually quoted to 2 decimal places and the fine structure (“coupling”) can be meaningful even if as small as 1 Hz (i.e. 0.02 ppm).
In the SPECTRa-We’ve been looking at how we can preserve this valuable data – it comes out of the machine in digital form, but then it is often transcribed into a PDF. Sometimes this preserves the graphics structure, sometimes it converts it to a pixellated image. This is the worst sort of hamburger.
Since the spectra are important tools in ensuring reproducibility, and chemists frequently refer to literature values, why do some journals allow such awful spectra. I suppose it’s better than having no spectra at all. Here are some good bad and ugly from supplemental info for recent synthetic chemistry papers. Since at least 3 of them carry a copyright I shan’t identify the journals. I claim that they are (a) data (b) a small portion of the work (c) publication does not affect sales (d) that most people would be ashamed to copyright them anyway.
Note that they all cover about 1 ppm (although for some you have to take the numerals on trust)
Fig. 1 The fuzz is real, but quite a bit is visible
Fig.2 Good. this seems to have preserved most of the data
Fig. 3 What are those figures??? Yes, I can guess – but I shouldn’t have to. But the limited pixel resolution has destroyed the peak shapes as well. Look at the non-linearity of the horizontal axis.
Fig 4. I’ve made this larger so the fuzziness from the pixellation is revealed.
Fig. 5 Quite good. You can certainly see peaks separated by 1 Hz.
Fig 6. Oh dear. This has the added fun of being a JPG which adds some dots to the spectrum which are nothing to do with the data. JPGs should not be used for this sort of thing.
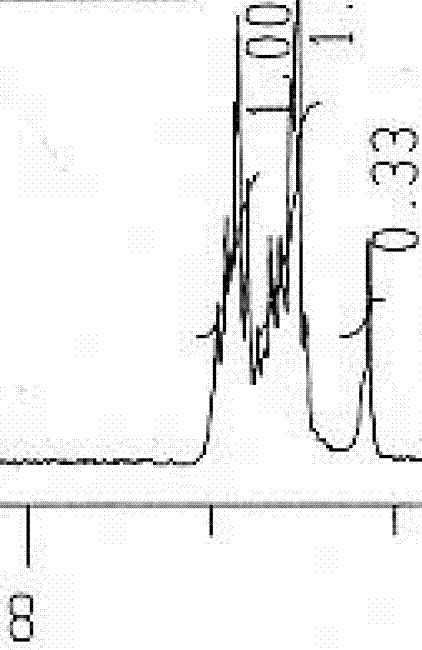
Fig 7. This is 8-7 ppm. Another JPG
So non-chemists should be able to see the point. If an article costs USD 3000 then the scientific community deserves better. How many chemists have cursed the unreadability of numeric data mangled by graphics tools? There is no technical reason why the digital data shouldn’t be deposited with the publisher, the instituion, the department.
The simple question is: do chemists care?
Peter – we certainly care deeply about the issue of NMR data loss.
There is thread on this going on in the UsefulChem mailing list right now:
http://groups.google.com/group/usefulchem/browse_thread/thread/ca5f52ce8ed44c2a
We’ve started uploading the JCAMP files of our starting materials and products into ChemSpider.
The spectra show up embedded in the molecule record using Robert Lancashire’s JSpecView. No information is lost – the spectra can be expanded and integrated at will and almost all the meta-data is automatically saved (solvent, aquisition time, etc.)
(1) J-C, of course my comments were not aimed at you but at the authors, editors and readers of journals – and I hope some of them comment.
I know the topic was Open data and it is good to see real spectra being stored. But how are the data being stored in ChemSpider and how available are they? Last time I looked the metadata was proprietary and the data were closed (there was a limit of – I think – 100 downloads).
I appreciate that not all the spectra are NMR, but have you considered putting those in NMRShiftDB. Then they would belong to the community.
I have plans which I hope to announce soon for an Open collection of spectra. No timescale.