Nick Day’s CrystalEye system can e thought of as an open, robotically managed, robotically quality-reviewed, data, overlay, “journal”. It’s not a conventional journal, but it ticks most of the buttons. And it publishes a new set of information each day. So I have subscribed to some of the RSS feeds on the site. There are many hundred – you can be alerted by journal, by chemical category, by bond type, by quality, etc. And they come naturally into the Feedreader. Here’s one from today. (It caught my eye because I worked on copper N,O chelates during my doctorate and it’s one of my favourite elemnts. Here’s what the feed delivered. No frills, no adverts, no javascript. Just simple science:

and here’s what you get if you follow the link:
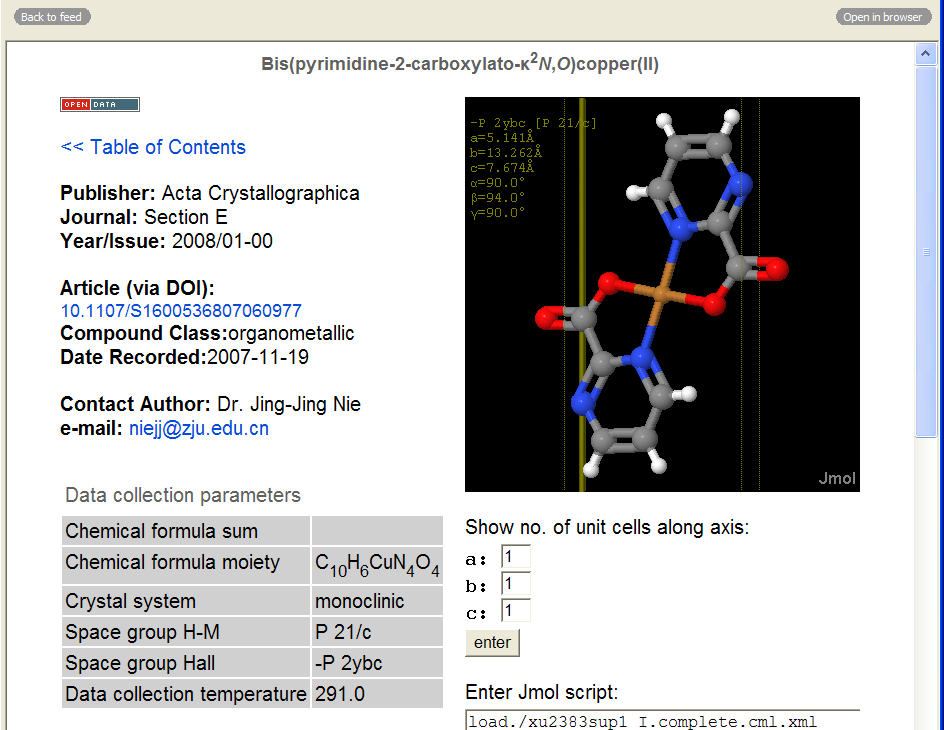
Click to enlarge.
This is immediate – Nick’s robots determine when a new issue has come out and various publishers are talking to us about providing RSS feeds of new issues or new articles (talked with BMC and IUCr yesterday). It makes it a lot simpler.
The possibilities are enormous. All the information is semantic and can be turned into RDF. Andrew Walkingshaw has done this and in a later post he or I will show how to search for information contained in the CIF files. If you are only interested in Cu-N bonds there is a special feed exactly for that purpose.
Interesting choice for the structure, Peter! 🙂
See: http://chem-bla-ics.blogspot.com/2008/01/why-chemistry-rich-rss-feeds-matter.html
(2) Egon = it was literlly the first one that emerged from the feed.
I will try to find some time this weekend to set up an agent to monitor the (CML)RSS feed to detect problems, which will be published as (CML)RSS again.
I receive the CrystalEye feeds via RSS. I find that >80% of them say:
No title supplied
from CrystalEye: All Structures by Chris Talbot
Is there a reason for this? I am assuming that the titles cannot be extracted?
Also, you might want to take a look at whether something has changed with CrystalEye of late. I’ve spent some time this week looking at the data sine we have been scraping. CrystalEye appears to be crashing both IE and firefox (it didn’t used to). It’s not unique to my single computer. I’ve tried it on three different computers.
Out of interest, what application/utility do you use to generate SMILES and InChI?
(3) I can’t help with (a) – there are a lot of structures that do not come from journals and those would not have titles. AFAIK Nick extracts titles from journal TOCs. But many of these TOCs are verbose HTML with styles and it may be difficult to find the title. (Journals with RSS TOCs avoid this problem). But I will let Nick answer.
(b) Can you be more specific? is it the RSS feed that crashes it, the searches, loading the indivdual pages, the applets or whatever. Suggest you mail Nick.
I do know that some versions of both browsers struggle with large tables and large HTML files. I think if the RSS was many thousands or the search returned many thousands that could crash some browsers.
(c) I suspect we use the SMILES tool in JUMBO. If not it would be OpenBabel. The InChI is generated with the standard InChI tool – there isn’t any option. Howe we wrap it varies but that is just part of the problem of managing C code in a Java environment. I am assuming the input is CML but Nick can say.
P
The first problem is due to the fact that CrystalEye uses the _publ_section_title field from the CIF for the titles. Initially, when it was just Acta and the RSC that we were scraping, virtually every CIF came with this field filled in. However, now a large percentage of the CIFs we get are from ACS and the COD, which seem to almost never have the title supplied. I shall alter the spiders so that the title is obtained from the TOC.
I’d be very interested to hear about any problems you’ve had with CrystalEye crashing browsers. Is it something that happens immediately, or after viewing a few structures?
I use Sam Adam’s jni-InChI library (on Sourceforge) to create the InChIs. It provides an easy way of creating InChIs from in-memory CML molecules in Java. The SMILES are generated using the CDK.
cheers,
Nick
I’ll be blogging shortly about our efforts to scrape CrystalEye as you suggested we should. I’ve come to a decision about what we will do to connect back to CrystalEye.
I believe you have a CML to SMILES conversion problem and will discuss how this observation helped make the decision.
Can I suggest that it would be of value to put a “Send Feedback” link on a record as we do with Curate Data on ChemSpider. Then I could send you/Nick/whoever commentaries when a record is seen to have errors. It doesn’t force an email exchange but rather is available for everyone to see. If someone sees an error then posts a comment it can be available for everyone and can therefore reduce mistakes. Just a thought. We’ve done it and have received hundreds of comments (http://www.chemspider.com/feedbackcurated.aspx) and made appropriate edits (not all yet!) This approach does not mean that you have to staff up to make corrections..but it does mean that the records are annotated to prevent other people making the same mistakes with the data (potentially) and does give you a way to find bugs etc.
(7) Thanks
The new data should be tractable through RSS. It is more effort, but we believe is tractable, to do the legacy data.
The spiders have now been altered to grab the titles from the TOCs rather than rely on them being in the CIF. It’s wasn’t difficult to do this (the TOCs are programmatically generated so the structure is repetitive) though the HTML structure of the pages are worthy of the ‘tagsoup’ label and so the XPaths required are ugly and usually have no semantics whatsoever.
Anyway, that should be the last of the ‘No title supplied’ message.
(9)Many thanks Nick. CrystalEye is much appreciated over here and will be an important part of the eChemistry project